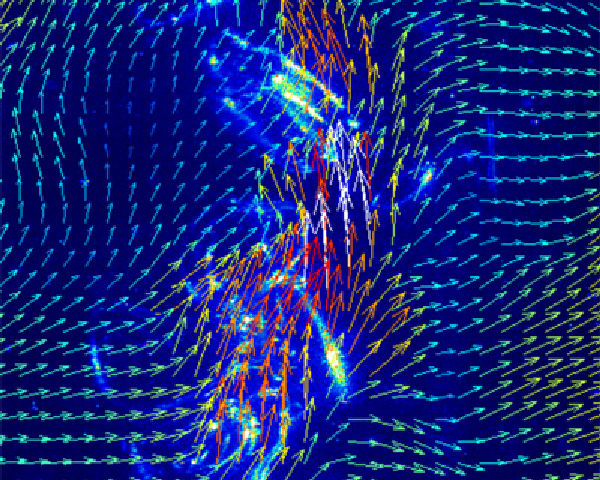

Dynamics of neural networks
The tools of dynamical systems theory are having an increasing impact on our understanding of patterns of neural activity. Some of the more popular single neuron models will be introduced and their behaviour in terms of bifurcation diagrams, phase-planes and phase-response curves will be explained.
For limit cycle oscillators, the coupled oscillator approach that has provided a framework for understanding behaviour in neural networks with weak synaptic and gap junction coupling will be reviewed. Then it will be shown how results for strong coupling can be obtained by focusing on a specific class of spiking neural models, namely (non-smooth) planar integrate-and-fire models.
It will be described how to build tractable tissue level models that maintain a strong link with biophysical reality. These models typically take the form of nonlinear integro-differential equations. Their non-local nature has led to the development of a set of analytical and numerical tools for the study of waves, bumps and patterns, based around natural extensions of those used for local differential equation models.
The relevance of neural field models for describing the brain at the large scales necessary for interpreting EEG and MEG data will be discussed. Recent results on next generation neural field models obtained via a mean field reduction from networks of nonlinear integrate-and-fire neurons will also be presented.
Meanfield theory for networks of spiking neurons
If functional units in the brain are large networks then we may not need to know the detailed dynamics of each neuron, but rather just a measure of the mean activity. Can we develop such a meanfield theory for networks of model spiking neurons?
The by now classical meanfield theory for irregularly spiking neurons will be discussed, which is based on two main assumptions: 1 - neurons receive large numbers of weak, uncorrelated inputs and 2 - neurons are Poisson processes. This is a very general theory which works for many spiking neuron models, but which only allows us to calculate stationary states, their stability, and the response to weak inputs.
A complementary theory will also be discussed, which only works for quadratic integrate-and-fire neurons with quenched variability, but which allows us to derive exact equations for the mean-field. These equations capture the full nonlinear response of the network and is simple enough to allow for extensive analysis.
Modeling interacting networks as processes with variable length
A class of recently introduced models to describe networks of neurons as stochastic processes with memory of variable length will be presented. These are non-Markovian processes in high or infinite dimension in which the past dependence of transition probabilities or intensities has a range that is finite but depends on the particular history.
Starting from existence results, we study related mean-field models in continuous time and their large population limits, and discuss the relation with associated Piecewise Deterministic Markov Processes (PDMP’s) and state results concerning their longtime behavior.
We will look at two important problems of statistical inference in such models: estimation of the spiking rate function and estimation of the neuronal interaction graph.

Detection of dependence between neurons and synchronisation
Synchronisation is an important phenomenon in neuroscience, and this kind of dependency between neuronal activity may play an important role for how the brain encodes information. An essential mathematical/statistical question is therefore how to detect such phenomena in a setup where the observations are scarce, noisy and always fluctuating.
The Unitary Events methods will be described (first introduced by Grün in the 90's) which makes rigorous the distinction between synchronization and pure coincidence and how statistical testing procedures can distinguish between them.
The notion of local independence between the spiking activity of neurons and how estimating a graph of local independence can infer functional connectivity will be explained.